Infectious agents – bacteria, viruses and parasites – evolve in response to selective pressures created by our immune systems, vaccine deployment, and antimicrobial drugs. The resulting genetic changes in the pathogens often have direct consequences for human health – most prominently, antibiotic resistance, but also ongoing transmission of pathogens that have evolved to evade our natural or vaccine-induced immune responses. Understanding the mechanisms by which this selection happens offers the opportunity to control it and direct it in more benign ways. Quantifying the contribution of particular evolutionary changes to disease burden allows priority-setting for antimicrobial stewardship, vaccine deployment and other responses. Evolutionary changes in pathogen genome sequences can also be used to track transmission and inform our understanding of who infects whom. Our research in this area ranges from very basic studies of mechanisms of natural selection in pathogens through applied efforts to direct evolution in ways that will improve human health.
Microbial evolution
Malaria genetics
The malaria parasite has fourteen chromosomes and undergoes sexual reproduction in its mosquito host. It exhibits extraordinary diversity on a population level, and this has hindered attempts to define “strains” to track transmission chains, as well as making it difficult to use genetic signatures to trace relatedness between parasite populations. Nevertheless, genetic signatures of evolutionary processes relevant for transmission (e.g. spatial spread) and selection (e.g. due to drug pressure), may provide valuable insights for control programs aiming to reduce and ultimately eliminate malaria. Currently, however, because of the complexities of malaria parasite biology, the methods required to make sense of the structure of malaria parasite populations – in terms of sequencing approaches, sampling strategies, and analytical tools – are still in their infancy. We are actively working to develop methods to analyze malaria parasite population genetic signatures, with a focus on tools to measure the spatial dynamics of malaria caused by human mobility in the context of national control program decision making.
Key publications
Quantifying connectivity between local Plasmodium falciparum malaria parasite populations using identity by descent. PLoS Genet. 2017
THE REAL McCOIL: A method for the concurrent estimation of the complexity of infection and SNP allele frequency for malaria parasites. PLoS Comput Biol. 2017
Variation in infection length and superinfection enhance selection efficiency in the human malaria parasite. Sci Rep. 2016
Ape parasite origins of human malaria virulence genes. Nat Commun. 2015
A network approach to analyzing highly recombinant malaria parasite genes. PLoS Comput Biol. 2013
Genomic epidemiology (LRE, pertussis, TB, Streptococcus pneumoniae)
As it has become easier and cheaper to determine the genome sequence of a pathogen, epidemiologists have increasingly made use of it to examine outbreaks, transmission patterns, and track the spread of individual strains of disease worldwide. CCDD has a long history of work using genomes to study the epidemiology of the pneumococcus, but has also pioneered the study of other bacterial pathogens including the agents that cause whooping cough, tuberculosis and gonorrhea as well as cases of the so-called “nightmare” drug resistant infections caused by Carbapenem resistant Enterobacteriaceae.
Key publications
Genomic epidemiology of Neisseria gonorrhoeae with reduced susceptibility to cefixime in the USA: a retrospective observational study. Lancet Infect Dis. 2014
Genomic epidemiology of the Escherichia coli O104:H4 outbreaks in Europe, 2011. Proc Natl Acad Sci U S A. 2012
Multi-institute analysis of carbapenem resistance reveals remarkable diversity, unexplained mechanisms, and limited clonal outbreaks. Proc Natl Acad Sci U S A. 2017
Bacterial genomes in epidemiology–present and future. Philos Trans R Soc Lond B Biol Sci. 2013
Rapid pneumococcal evolution in response to clinical interventions. Science. 2011
Population genomics of post-vaccine changes in pneumococcal epidemiology. Nat Genet. 2013
Selection by natural immunity
Evolution of Streptococcus pneumoniae in response to the human immune system

Human hosts gradually become immune to carriage of and disease from Streptococcus pneumoniae, developing both antibody responses to specific serotypes, and antibody and cellular (Th17) responses to protein antigens on the pneumococcus. Over more than a decade, we have characterized the specificity of these immune responses, their effects on patterns of carriage as individuals age, the way they structure the pneumococcal population, and the consequences for patterns of human disease
Key publications
The impact of serotype-specific vaccination on phylodynamic parameters of Streptococcus pneumoniae and the pneumococcal pan-genome. PLoS Pathog. 2018
Frequency-dependent selection in vaccine-associated pneumococcal population dynamics. Nat Ecol Evol. 2017
Diverse evolutionary patterns of pneumococcal antigens identified by pangenome-wide immunological screening. Proc Natl Acad Sci U S A. 2017
Association of Pneumococcal Protein Antigen Serology With Age and Antigenic Profile of Colonizing Isolates. J Infect Dis. 2017
Effect of Serotype on Pneumococcal Competition in a Mouse Colonization Model. MBio. 2015
Selective and genetic constraints on pneumococcal serotype switching. PLoS Genet. 2015
Population genomics of post-vaccine changes in pneumococcal epidemiology. Nat Genet. 2013
Distinct effects on diversifying selection by two mechanisms of immunity against Streptococcus pneumoniae. PLoS Pathog. 2012
Niche and neutral effects of acquired immunity permit coexistence of pneumococcal serotypes. Science. 2012
Pneumococcal capsular polysaccharide structure predicts serotype prevalence. PLoS Pathog. 2009
Interleukin-17A mediates acquired immunity to pneumococcal colonization. PLoS Pathog. 2008
Selection by vaccines
Selection by vaccines
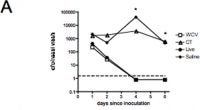
Medical interventions like antibiotics and vaccines are novel selective pressures that pathogens must face, and we can learn a lot from how they respond to them. Some vaccines either in use or in development target the most dangerous known strains of a pathogen, which might remove competition that has been keeping other strains rare, leading these rare strains to become more common and increased exposure. If these emerging strains are either more virulent than we thought, or become more virulent, the effectiveness of the vaccines might be reduced.
Researchers in CCDD have been studying this in multiple pathogens, but with a special focus on the pneumococcus.
Key publications
Population genomics of post-vaccine changes in pneumococcal epidemiology. Nat Genet. 2013
Stability of the pneumococcal population structure in Massachusetts as PCV13 was introduced. BMC Infect Dis. 2015
Interim results of an ecological experiment – Conjugate vaccination against the pneumococcus and serotype replacement. Hum Vaccin Immunother. 2016
Serotype replacement in disease after pneumococcal vaccination. Lancet. 2011
Population genetic structure, antibiotic resistance, capsule switching and evolution of invasive pneumococci before conjugate vaccination in Malawi. Vaccine. 2017
Genomic Epidemiology of Penicillin-Nonsusceptible Pneumococci with Nonvaccine Serotypes Causing Invasive Disease in the United States. J Clin Microbiol. 2017
Vaccines as a solution to AMR
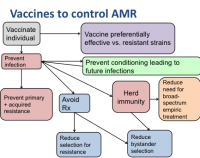
Vaccines are an underappreciated part of the solution to the growing problem of antimicrobial resistance in bacteria. By preventing bacterial infections with bacteria, vaccines such as the pneumococcal conjugate vaccines reduce the burden of resistant as well as non-resistant infections. Moreover, by preventing infections that can lead to antimicrobial use, vaccines against both bacteria and other pathogens (eg influenza) help to reduce the intensity of selection on bacteria to become resistant.
CCDD researchers are working to delineate the mechanisms by which vaccines can reduce AMR and quantify the contribution of various mechanisms. Also see AMR.
Key publications
Pan-serotype Reduction in Progression of Streptococcus pneumoniae to Otitis Media After Rollout of Pneumococcal Conjugate Vaccines. Clin Infect Dis. 2017
How Can Vaccines Contribute to Solving the Antimicrobial Resistance Problem? MBio. 2016
Targeting imperfect vaccines against drug-resistance determinants: a strategy for countering the rise of drug resistance. PLoS One. 2013
Estimating the proportion of bystander selection for antibiotic resistance in the US. Preprint. 2018
Antimicrobial resistance
AMR
Antimicrobial resistance is a growing concern in the developed and the developing world. Strains of several bacteria untreatable with any standard drug have been observed, and the narrowing therapeutic options threaten not only the ability to treat infections but the feasibility of surgeries and cancer chemotherapies that rely on antibiotic prophylaxis to prevent infection.
CCDD researchers are tackling multiple aspects of this problem, including: 
• Advancing the basic science of how antibiotic use is related to antibiotic resistance
• Using genome sequences to study the spread of resistant bacteria
• Developing new diagnostic predictors for antibiotic resistance
• Prioritizing interventions to slow and reverse the rise of resistance
• Critically assessing the role of agricultural antibiotic use in driving resistance
• Understanding the population drivers of multiple drug resistance
Key publications
Mechanisms that maintain coexistence of antibiotic sensitivity and resistance also promote high frequencies of multidrug resistance. Preprint. 2017
Evolution of antibiotic resistance is linked to any genetic mechanism affecting bacterial duration of carriage. Proc Natl Acad Sci U S A. 2017
Host population structure and treatment frequency maintain balancing selection on drug resistance. J R Soc Interface. 2017
Multi-institute analysis of carbapenem resistance reveals remarkable diversity, unexplained mechanisms, and limited clonal outbreaks. Proc Natl Acad Sci U S A. 2017
Genomic Epidemiology of Gonococcal Resistance to Extended-Spectrum Cephalosporins, Macrolides, and Fluoroquinolones in the United States, 2000-2013. J Infect Dis. 2016
Geographic and temporal trends in antimicrobial nonsusceptibility in Streptococcus pneumoniae in the post-vaccine era in the United States. J Infect Dis. 2013
Antibiotics in agriculture and the risk to human health: how worried should we be? Evol Appl. 2015
Factors related to increasing prevalence of resistance to ciprofloxacin and other antimicrobial drugs in Neisseria gonorrhoeae, United States. Emerg Infect Dis. 2012
Incremental increase in fitness cost with increased beta -lactam resistance in pneumococci evaluated by competition in an infant rat nasal colonization model. J Infect Dis. 2006
Geographic diversity and temporal trends of antimicrobial resistance in Streptococcus pneumoniae in the United States. Nat Med. 2003
Measuring and interpreting associations between antibiotic use and penicillin resistance in Streptococcus pneumoniae. Clin Infect Dis. 2001
Antimicrobial use and antimicrobial resistance: a population perspective. Emerg Infect Dis. 2002
Vaccines as solution to AMR
Cross-listed with Selection by vaccines research subcategory
Harnessing evolution
Detecting transmission
Cross-listed with Predicting and Detecting Infectious Disease research category–> Surveillance methods–> Detecting transmission
Rapid detection of antimicrobial resistance (AMR)
Cross-listed with Predicting and Detecting Infectious Disease research category–> Surveillance methods–> Rapid detection of AMR
Enhancing vaccine trials with pathogen sequence data
Cross-listed with Quantifying and Controlling Infectious Disease research category–>Clinical trial preparedness–>Enhancing vaccine trials with new data sources
Engineering pathogen diversity to study population dynamics within hosts and the distribution of host susceptibility
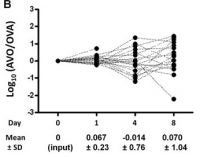
More than one strain of a pathogen may infect a particular host simultaneously, and population bottlenecks during that process may have consequences for pathogen evolution. Using experimental infections with bacterial and viral pathogens, CCDD researchers study the patterns of pathogen diversity as an infection establishes and progresses, thereby making inferences about the infectious process.
Key publications
Vaccine Effects on Heterogeneity in Susceptibility and Implications for Population Health Management. MBio. 2017
Sequence tag-based analysis of microbial population dynamics. Nat Methods. 2015
Within-host selection is limited by an effective population of Streptococcus pneumoniae during nasopharyngeal colonization. Infect Immun. 2013