In an era when an infected person or vector can cross the world in a day, preventing and containing outbreaks of disease anywhere protects the health of people everywhere. Every large infectious disease threat begins as one or a few cases and becomes widespread only because containment efforts were not tried or did not succeed. Our research enhances preparedness to enable rapid, rational and effective countermeasures to control the spread of disease in new outbreaks and in new settings, driven by urbanization and other forms of human mobility.
Surveillance methods
Rapid detection of antimicrobial resistance
Drug resistant infections are a cause of growing concern, complicating treatment of potentially lethal disease and important causes of morbidity, mortality and increased healthcare costs. In the US it is estimated that a person dies of a drug resistant infection roughly every twenty minutes. The sooner effective therapy can be started the better, but how can we know which bacterium is causing disease and whether it is resistant?
Researchers in CCDD are exploring the use of new technologies capable of sequencing in ‘real time’ using a USB connected portable sequencer that runs off a standard laptop. As the sequence data are generated, they can be checked against a database to determine whether sequence characteristic of a resistant infection is present, and providing useful information more quickly – on a time scale of minutes.
Detecting transmission
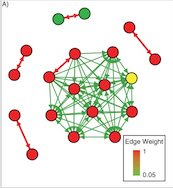 Transmission is central to infectious disease epidemiology, being both the first step in the path to illness and the key determinant of epidemic dynamics. Knowing the complete transmission tree, ie exactly how all the cases of disease are linked together, is useful for estimating things like the basic reproductive number, the who-infected-whom matrix, and would be invaluable in outbreak investigations.
Transmission is central to infectious disease epidemiology, being both the first step in the path to illness and the key determinant of epidemic dynamics. Knowing the complete transmission tree, ie exactly how all the cases of disease are linked together, is useful for estimating things like the basic reproductive number, the who-infected-whom matrix, and would be invaluable in outbreak investigations.
Researchers in CCDD are actively working to develop and apply methods for detecting transmission that make use of genomic sequence data. Because DNA sequencing is increasingly routine this is potentially a rich seam of information. The simple principle is that the genomes of pathogens from linked cases should be more similar than those from unlinked cases. However this assumes that a case of disease may be characterized easily, with one genome chosen to represent what could be considerable variation within the pathogen population. Because of the relatively high rate with which pathogens can mutate, this within host diversity can be great, and we are investigating ways in which it might be an additional source of variation to link cases. We are also working with simulations of pathogen evolution to provide guidelines for doctors who want to know whether a new case of disease has been imported from elsewhere, or is related to something that has been already sampled.
Key publication
Shared Genomic Variants: Identification of Transmission Routes Using Pathogen Deep-Sequence Data. Am J Epidemiol. 2017
Expository Reviews
Productive disruption: opportunities and challenges for innovation in infectious disease surveillance. BMJ Global Health.2018.
Mapping malaria by combining parasite genomic and epidemiologic data.BMC Medicine.2018
Nowcasting
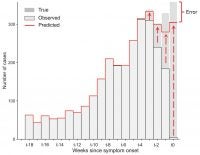
Infectious disease data are reported to authorities, such as health departments or the CDC, over time, with delay from the time a patient gets sick to the time the case is reported. Accounting for such reporting delays is essential for authorities to have a clear picture of the current state of an epidemic, as many recent cases will not yet have been reported, leading to an apparent decline of cases toward the present. We are working on methods for Bayesian nowcasting, to learn about the pattern of delays from past weeks in order to estimate what proportion of recent cases have yet to be reported, and thus estimate the number of current cases.
Alert thresholds
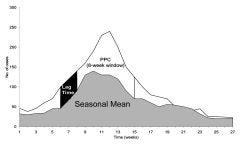
Disease control activities are costly and disruptive, so they should ideally be used only when they can prevent substantial numbers of cases. Such interventions can be triggered when the number of cases or some other indicator of disease activity passes an “alert threshold” that indicates unusually high activity. Such triggers should be sensitive (picking up real outbreaks), specific (not triggering unless a real outbreak is in progress) and timely (giving ample warning and time to implement control measures). We have developed measures to express the optimal combination of these characteristics in a disease- and control-measure-specific way, thereby providing a single method for comparing alert thresholds. Originally developed for malaria control, these methods are being adapted for hospital infection control interventions.
Key publication
Alert threshold algorithms and malaria epidemic detection. Emerg Infect Dis. 2004
Seasonality
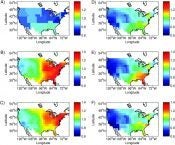
Since Thucydides it has been known that infectious diseases vary seasonally. We have elucidated the causes of seasonality in influenza, dissecting the contributions of school calendars (important), humidity-dependent virus survival (important), and varying levels of vitamin D (less important). More recently, we have focused on seasonality in other infections.
Human mobility
Human travel is one of the most important drivers of infectious disease outbreaks; infected travelers introduce infections to susceptible populations and can spread disease on different spatial scales. Until recently, however, for most places – particularly low and middle income settings – it has been almost impossible to find robust, scalable data on this key component of epidemic spread. With the rapid adoption of mobile phones across the globe, mobile operators are now collecting vast volumes of real-time information about the location and dynamics of human populations at scale. Researchers at the center are pioneering the use of these data to model a) the spatial spread of malaria, dengue, and other infectious diseases, b) the response of human populations to natural disasters like flooding, and c) the impact of travel restrictions and reactive vaccination campaigns on the containment of emerging epidemics. We are also working closely with operators to develop industry-wide rules for anonymization of these data sets that protect the privacy of subscribers.
Quantifying the impact of human mobility on malaria. Science. 2012
Policy
Cholera vaccines
The efficacy of containment strategies deployed in the case of an emerging epidemic threat will depend on the epidemiological features of the pathogen in question, as well as availability of pharmaceutical interventions, if any. At CCDD we have been developing theoretical frameworks for decision making with respect to quarantine and symptom monitoring, in the case of epidemics that do not have pharmaceutical interventions, and with respect to vaccination roll-out in the case of epidemic cholera. These models combine the dynamics of pathogen transmission with the timing of interventions (e.g. the timing of vaccination roll-out and dosing strategies) will be key to effective containment.
Key publications
Prolonging herd immunity to cholera via vaccination: Accounting for human mobility and waning vaccine effects. PLoS Negl Trop Dis. 2018
Science policy: potential pandemic pathogens
 CCDD Director Dr. Marc Lipsitch has written extensively on the risks and scientific limitations of experiments to create “potential pandemic pathogens” (PPP) by modifying influenza viruses to be readily transmissible among mammals (ferrets, the model for human infection).
CCDD Director Dr. Marc Lipsitch has written extensively on the risks and scientific limitations of experiments to create “potential pandemic pathogens” (PPP) by modifying influenza viruses to be readily transmissible among mammals (ferrets, the model for human infection).
Key publications
Commentary: Six policy options for conducting gain-of-function research. CIDRAP. 2016
Moratorium on research intended to create novel potential pandemic pathogens. MBio. 2014
Science policy: preprints
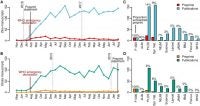 CCDD researchers have been strong advocates and users of preprints to accelerate communication and progress during outbreaks of infectious diseases, as well as to enhance scientific openness in general.
CCDD researchers have been strong advocates and users of preprints to accelerate communication and progress during outbreaks of infectious diseases, as well as to enhance scientific openness in general.
Key publications
Preprints: An underutilized mechanism to accelerate outbreak science. PLoS Med. 2018
Ethics
Ethics and infectious diseases
Infectious diseases raise special concerns in research ethics. These arise both from the scale of epidemics that can threaten populations, from the resulting urgency and logistical and political complexity of doing trials, and from the fact that research seeks to understand not only the protection a vaccine or treatment can offer an individual, but often also what it can offer that individual’s contacts in the population. CCDD researchers work at the intersection of study design, mathematical modeling, and research ethics to define ethical, scientifically valid studies in crisis situations and to alert the scientific community to the hazards of especially dangerous laboratory research.
Key publications
Vaccine testing for emerging infections: the case for individual randomisation. J Med Ethics. 2017
New Game, New Rules: Limiting the Risks of Biological Engineering. Foreign Affairs. 2015
Additional article
Underprotection of Unpredictable Statistical Lives Compared to Predictable Ones. Risk Anal. 2017